CLASHub Database Tutorial
This short tutorial walks you through the three database search modes available on the CLASHub home page: CLASH, Gene Expression, and microRNA Expression. You’ll learn the required steps, how autocomplete works, and how to avoid common submission issues.
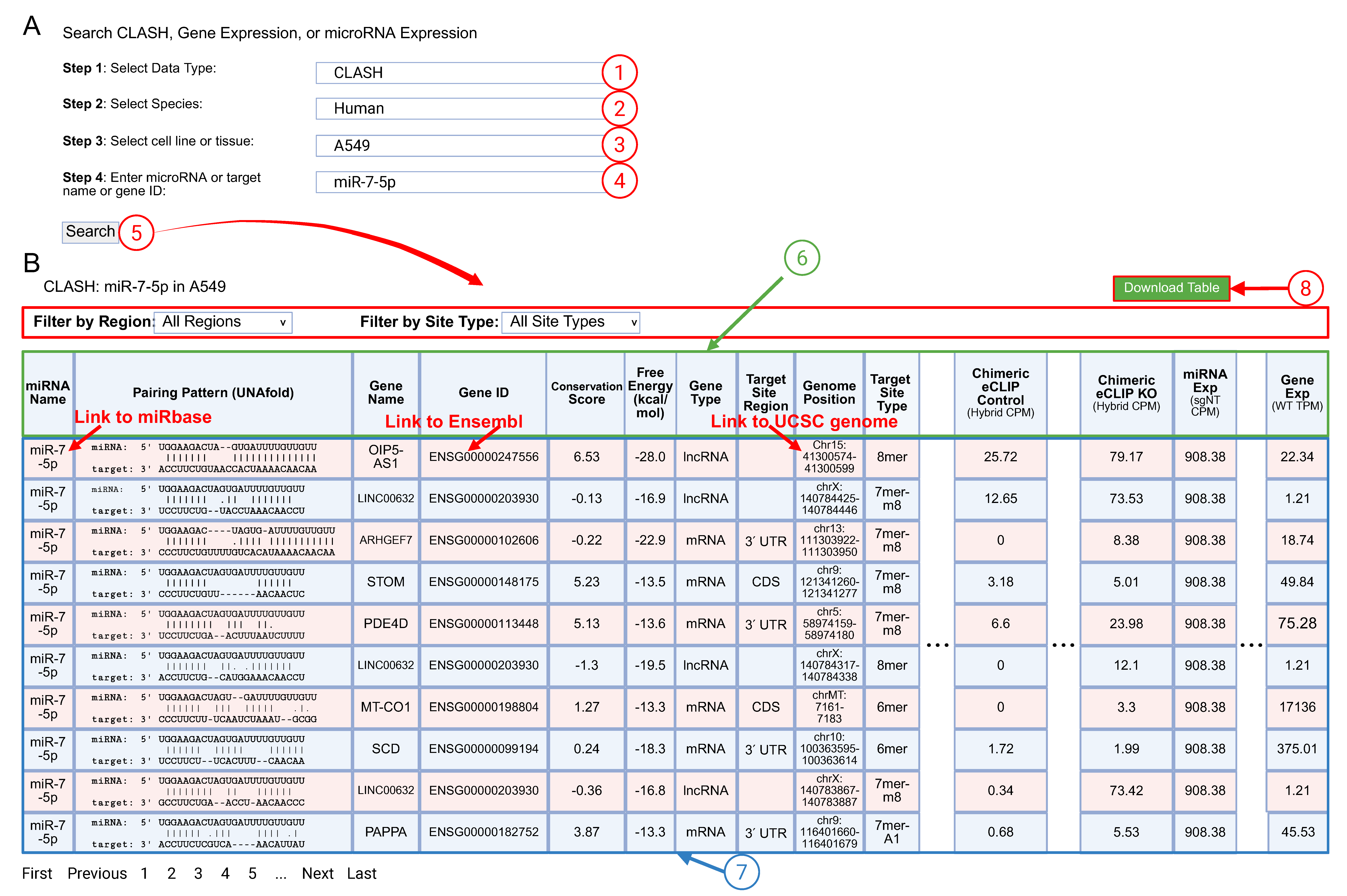
(A) The CLASHub web interface guides users by selecting 1 data type, 2 species, 3 sample type and 4 miRNA or gene ID in the query fields, then performing the 5 Search function.
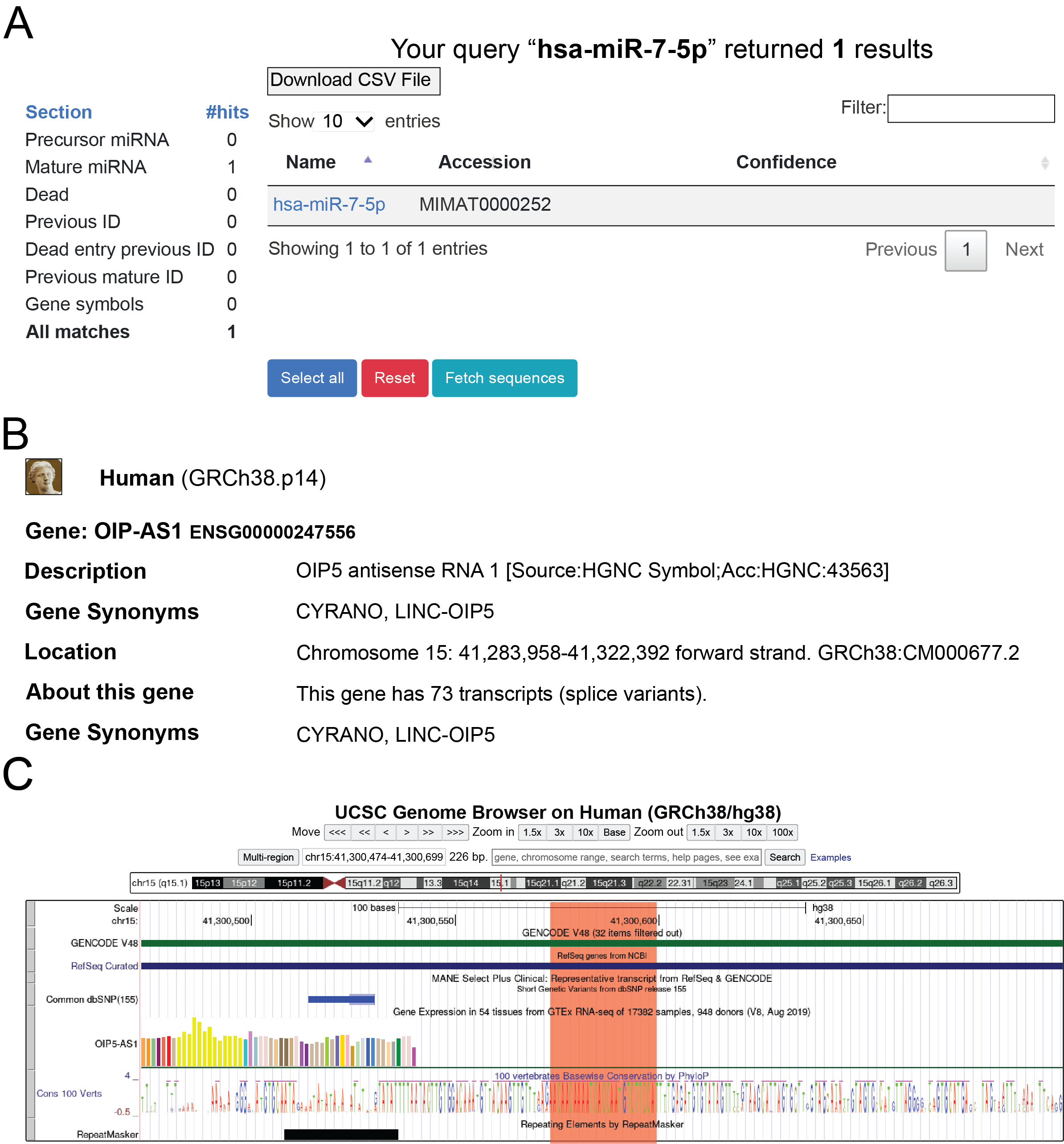
(B) The example CLASH output table summarizes detected miR‑7 hybrids in A549 cells (6). Each row (7) includes comprehensive information (miRNA name, predicted pairing pattern, gene, gene ID, genome position, target site type, and abundance metrics). The 8 Download Table button exports the table for offline analysis. Interactive links allow opening miRBase (via miRNA name), Ensembl (via gene ID), and UCSC Genome Browser (via genome position).
External resources
Gene & microRNA Expression (RNA/miRNA-seq)